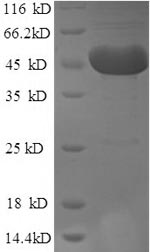
m7GpppN-mRNA hydrolase (DCP2) is a crucial enzyme involved in mRNA decay processes. DCP2, a conserved NUDIX hydrolase, functions as a decapping enzyme by cleaving the 5’ m7G cap of mRNA, leading to the release of m7GDP and a 5’-end monophosphorylated RNA [1][2][3]. It has been shown to be an essential component of the decapping complex and is involved in the hydrolysis of cap structures generated by mRNA decay [4][2]. DCP2 is an RNA-binding protein that selectively binds to specific mRNAs containing a stem-loop structure near the 5' cap, indicating its role in selective mRNA decay [5]. Additionally, DCP2 has been implicated in various mRNA decay processes, suggesting its differential utilization in different pathways [6]. The enzyme's activity can be influenced by both sequence and context, indicating its potential contribution to gene regulation in multiple RNA pathways [7].
Furthermore, DCP2 is a central player in mRNA metabolism, and its molecular basis of mRNA decapping has been intensively studied [8]. It is conserved among eukaryotes and is considered the major decapping enzyme in yeast, indicating its significance in the 5′ to 3′ mRNA decay pathway [9][10]. DCP2 has also been shown to interact with other proteins, such as tristetraprolin (TTP), to mediate mRNA decay, highlighting its role in facilitating the assembly of decay-competent mRNPs [11]. Moreover, DCP2's function is not limited to yeast, as it has been implicated in mRNA decay processes in mammalian cells, raising questions about its exclusivity as the major decapping enzyme in multicellular organisms [12].
References:
[1] J. Lobel, R. Tibble, & J. Gross, "Pat1 activates late steps in mrna decay by multiple mechanisms",, 2019. https://doi.org/10.1101/594168
[2] S. Kramer, "The apah-like phosphatase tbalph1 is the major mrna decapping enzyme of trypanosomes", Plos Pathogens, vol. 13, no. 6, p. e1006456, 2017. https://doi.org/10.1371/journal.ppat.1006456
[3] C. Chang, N. Bercovich, B. Loh, S. Jonas, & E. Izaurralde, "The activation of the decapping enzyme dcp2 by dcp1 occurs on the edc4 scaffold and involves a conserved loop in dcp1", Nucleic Acids Research, vol. 42, no. 8, p. 5217-5233, 2014. https://doi.org/10.1093/nar/gku129
[4] V. Taverniti and B. Séraphin, "Elimination of cap structures generated by mrna decay involves the new scavenger mrna decapping enzyme aph1/fhit together with dcps", Nucleic Acids Research, vol. 43, no. 1, p. 482-492, 2014. https://doi.org/10.1093/nar/gku1251
[5] E. Grudzien-Nogalska and M. Kiledjian, "New insights into decapping enzymes and selective mrna decay", Wiley Interdisciplinary Reviews - Rna, vol. 8, no. 1, 2016. https://doi.org/10.1002/wrna.1379
[6] Y. Li, M. Song, & M. Kiledjian, "Differential utilization of decapping enzymes in mammalian mrna decay pathways", Rna, vol. 17, no. 3, p. 419-428, 2011. https://doi.org/10.1261/rna.2439811
[7] L. Cohen, C. Mikhli, X. Jiao, M. Kiledjian, G. Kunkel, & R. Davis, "Dcp2 decaps m2,2,7gpppn-capped rnas, and its activity is sequence and context dependent", Molecular and Cellular Biology, vol. 25, no. 20, p. 8779-8791, 2005. https://doi.org/10.1128/mcb.25.20.8779-8791.2005
[8] M. Deshmukh, B. Jones, D. Quang-Dang, J. Flinders, S. Floor, C. Kimet al., "Mrna decapping is promoted by an rna-binding channel in dcp2", Molecular Cell, vol. 29, no. 3, p. 324-336, 2008. https://doi.org/10.1016/j.molcel.2007.11.027
[9] M. Kim and A. Hoof, "Suppressors of mrna decapping defects isolated by experimental evolution ameliorate transcriptome disruption without restoring mrna decay",, 2020. https://doi.org/10.1101/2020.06.14.151068
[10] S. Erickson, E. Corpuz, J. Maloy, C. Fillman, W. K, E. Bennettet al., "Competition between decapping complex formation and ubiquitin-mediated proteasomal degradation controls human dcp2 decapping activity", Molecular and Cellular Biology, vol. 35, no. 12, p. 2144-2153, 2015. https://doi.org/10.1128/mcb.01517-14
[11] V. Maciej, N. Mateva, J. Schwarz, T. Dittmers, M. Mallick, H. Urlaubet al., "Intrinsically disordered regions of tristetraprolin and dcp2 directly interact to mediate decay of are-mrna", Nucleic Acids Research, vol. 50, no. 18, p. 10665-10679, 2022. https://doi.org/10.1093/nar/gkac797
[12] M. Song, L. You, & M. Kiledjian, "Multiple mrna decapping enzymes in mammalian cells", Molecular Cell, vol. 40, no. 3, p. 423-432, 2010. https://doi.org/10.1016/j.molcel.2010.10.010